rfp lab
purpose
The purpose of this lab was to make RFP (Red Fluorescent Protein) from the genes of jelly fish in bacteria and to learn the steps of genetic engineering.
materials & procedure
2a) Materials and procedure can be found in Amgen lab manual 2a.
4a) Materials and procedure can be found in Amgen lab manual 4a.
5a) Materials and procedure can be found in Amgen lab manual 5a.
6a) Materials and procedure can be found in Amgen lab manual 6.
4a) Materials and procedure can be found in Amgen lab manual 4a.
5a) Materials and procedure can be found in Amgen lab manual 5a.
6a) Materials and procedure can be found in Amgen lab manual 6.
Experimental Overview
Lab 2a:
Verification of a plasmid by restriction digest
-Cut plasmid with BanHI and HindIII to cut out RFP-ara from bacteria plasmid
Lab 4a:
Verification of plasma digest by gel electrophoresis
Lab 5a:
Transformation of bacteria with recombinant plasmid
Lab 6:
Portification of RFP using chromatography
Verification of a plasmid by restriction digest
-Cut plasmid with BanHI and HindIII to cut out RFP-ara from bacteria plasmid
Lab 4a:
Verification of plasma digest by gel electrophoresis
Lab 5a:
Transformation of bacteria with recombinant plasmid
Lab 6:
Portification of RFP using chromatography
in-lAB QUESTIONS
Lab 2a
Before 2a:
1. If pARA-R is digested with BamHI and HindIII, what fragments are produced? Record the nucleotide sequence of the sticky ends and the length of each fragment (bp), and indicate the genes and other important sequences present on each fragment.
-The two fragments produced are an RFP fragment with pBAD and an Ara-C fragment with ori and Amp-R. The nucleotide sequence length of the RFP fragment is 807 BP and the Ara-C fragment is 4495 BP.
2. In order to create a plasmid that can produce the red fluorescent protein in bacteria, what components are needed in the plasmid?
-To produce the red fluorescent protein in our bacteria, the RFP gene as well as Ara-C (which binds to the promoter region) are needed.
3. Bacteria can be killed by an antibiotic unless they carry a plasmid that has the gene for resistance to that antibiotic. Bio technologists call these genes selectable markers because only bacteria that carry the gene will survive an antibiotic. If the uptake of DNA by bacteria is inefficient, why is a selectable marker critical in cloning a gene in bacteria?
-A selectable marker is extremely important to cloning genes in bacteria because it determines which bacteria will continue to grow, while the bacteria without the marker die, thus separating the two colonies.
During 2a:
1. List in words or indicate in a drawing the important features of a plasmid vector that are required to clone a gene. Explain the purpose of each feature.
Plasmid Vector:
2. What role do restriction enzymes have in nature?
-Restriction enzymes act as a natural defense against bacteria. They cut out DNA of other bacteria (killing them) to help their own chances of survival.
3. Using your understanding of evolution, why would bacteria retain a gene that gives them resistance to antibiotics? How is the existence of bacteria with antibiotic resistance affecting medicine today?
-Antibiotic resistance genes would be kept because bacteria could survive in the presence of what could be lethal. These resistant bacteria affect medicine because stronger antibiotics are needed for less severe bacteria as well as new methods of killing bacteria needing to be developed for those strong bacteria.
4. Bacteria, sea anemones, and humans seem, on the surface, to be very different organisms. Explain how a gene from humans or a sea anemone can be expressed in bacteria to make a product never before made in bacteria.
-All genes have the same central dogma (DNA -> mRNA ->Protein), so they can be expressed in seemingly different organisms.
5. Due to a mishap in the lab, bacteria carrying a plasmid with an ampacillin resistant gene and bacteria carrying a plasmid with a gene that provides resistance to another antibiotic (kanamycin) were accidentally mixed together. Design an experiment that will allow you to sort out the two kinds of bacteria.
-Put half of the colony in a petri dish of ampicillin and the other half into a dish of kanamycin. The ampacillin dish will be left with only ampacillin-resistant bacteria and the kanamycin dish will only have kanamycin-resistant bacteria.
1. If pARA-R is digested with BamHI and HindIII, what fragments are produced? Record the nucleotide sequence of the sticky ends and the length of each fragment (bp), and indicate the genes and other important sequences present on each fragment.
-The two fragments produced are an RFP fragment with pBAD and an Ara-C fragment with ori and Amp-R. The nucleotide sequence length of the RFP fragment is 807 BP and the Ara-C fragment is 4495 BP.
2. In order to create a plasmid that can produce the red fluorescent protein in bacteria, what components are needed in the plasmid?
-To produce the red fluorescent protein in our bacteria, the RFP gene as well as Ara-C (which binds to the promoter region) are needed.
3. Bacteria can be killed by an antibiotic unless they carry a plasmid that has the gene for resistance to that antibiotic. Bio technologists call these genes selectable markers because only bacteria that carry the gene will survive an antibiotic. If the uptake of DNA by bacteria is inefficient, why is a selectable marker critical in cloning a gene in bacteria?
-A selectable marker is extremely important to cloning genes in bacteria because it determines which bacteria will continue to grow, while the bacteria without the marker die, thus separating the two colonies.
During 2a:
1. List in words or indicate in a drawing the important features of a plasmid vector that are required to clone a gene. Explain the purpose of each feature.
Plasmid Vector:
- Ori: Origin of replication
- RFP and PBad: Gene of interest
- Amp-R: Selective marker
- Ara-C: Binds to the promoter region which leads to transcription
2. What role do restriction enzymes have in nature?
-Restriction enzymes act as a natural defense against bacteria. They cut out DNA of other bacteria (killing them) to help their own chances of survival.
3. Using your understanding of evolution, why would bacteria retain a gene that gives them resistance to antibiotics? How is the existence of bacteria with antibiotic resistance affecting medicine today?
-Antibiotic resistance genes would be kept because bacteria could survive in the presence of what could be lethal. These resistant bacteria affect medicine because stronger antibiotics are needed for less severe bacteria as well as new methods of killing bacteria needing to be developed for those strong bacteria.
4. Bacteria, sea anemones, and humans seem, on the surface, to be very different organisms. Explain how a gene from humans or a sea anemone can be expressed in bacteria to make a product never before made in bacteria.
-All genes have the same central dogma (DNA -> mRNA ->Protein), so they can be expressed in seemingly different organisms.
5. Due to a mishap in the lab, bacteria carrying a plasmid with an ampacillin resistant gene and bacteria carrying a plasmid with a gene that provides resistance to another antibiotic (kanamycin) were accidentally mixed together. Design an experiment that will allow you to sort out the two kinds of bacteria.
-Put half of the colony in a petri dish of ampicillin and the other half into a dish of kanamycin. The ampacillin dish will be left with only ampacillin-resistant bacteria and the kanamycin dish will only have kanamycin-resistant bacteria.
lab 4a
Before 4a:
1. The pARA-R plasmid you digested in Laboratory 2A was replicated in a bacterial cell. What configurations—supercoiled, nicked circle, and multimer—might the plasmid have before digestion?
-The plasmid can have all three configurations.
1. The pARA-R plasmid you digested in Laboratory 2A was replicated in a bacterial cell. What configurations—supercoiled, nicked circle, and multimer—might the plasmid have before digestion?
-The plasmid can have all three configurations.
|
Tube
R+ R- |
Fragments (BP size smallest --> largest)
|
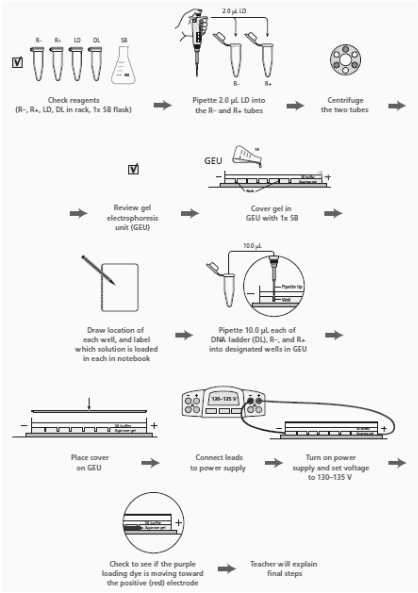
3. Read through the Methods section on pages 64 through 66 and briefly outline the steps, using words and a flowchart.
During 4a:
- Why is it important to verify that you have the correct recombinant plasmid?
-It is important to verify because during the lab, because we could make measuring mistakes, the materials could be faulty, or make errors in our procedure. - How did your actual gel results compare to your gel predictions?
-Our gel worked well except there were missing bands were the loading dye was meant to be seen that we did not predict.
3. Do you see any bands that are not expected? What could explain the origin of these unexpected bands?
-Our group had no unexpected bands. but we did not expect there to be missing bands.
4. Does the gel show that you are using the correct recombinant plasmid? Describe the evidence you used to make this assessment.
-We know that we are using the correct plasmid because the two bands that appear, and their correct positions.
5. In the R– lane, do you see evidence of multiple configurations of plasmids? Explain your answer.
-Yes, we know this because there are multiple bands and the different bands are different widths.
6. In the R+ lane, do you see evidence of complete digestion? Explain your answer.
-You can tell that complete digestion has occurred, because there are only two bands in the R+ lane.
7. In which lane would you expect to find the RFP gene and the ampR gene in the gel photograph? Are you able to locate these two genes? Explain your answer.
-In the R+ lane we would see a band around 4500 bp, and in the R- lane the band would be located at the 807 bp position. We can see this in the different lanes in the picture. The RFP gene would be in the R- lane, while the ampR gene would be in the R+ lane.
8. Compare the lanes that have linear fragments with the lanes that have plasmids. Is there a difference in the shape of the bands between these two DNA forms?
-Some bands have a wider shape. The lanes that have linear fragments have treading edges while the others have non treading edges.
-Our group had no unexpected bands. but we did not expect there to be missing bands.
4. Does the gel show that you are using the correct recombinant plasmid? Describe the evidence you used to make this assessment.
-We know that we are using the correct plasmid because the two bands that appear, and their correct positions.
5. In the R– lane, do you see evidence of multiple configurations of plasmids? Explain your answer.
-Yes, we know this because there are multiple bands and the different bands are different widths.
6. In the R+ lane, do you see evidence of complete digestion? Explain your answer.
-You can tell that complete digestion has occurred, because there are only two bands in the R+ lane.
7. In which lane would you expect to find the RFP gene and the ampR gene in the gel photograph? Are you able to locate these two genes? Explain your answer.
-In the R+ lane we would see a band around 4500 bp, and in the R- lane the band would be located at the 807 bp position. We can see this in the different lanes in the picture. The RFP gene would be in the R- lane, while the ampR gene would be in the R+ lane.
8. Compare the lanes that have linear fragments with the lanes that have plasmids. Is there a difference in the shape of the bands between these two DNA forms?
-Some bands have a wider shape. The lanes that have linear fragments have treading edges while the others have non treading edges.
lab 5a
Before 5a:
- Ampicillin is an antibiotic that kills bacterial cells by disrupting the formation of cell walls. However, the pARA-R plasmid has the ampicillin resistance gene, which produces a protein that breaks down ampicillin. What is the purpose of growing bacteria that have been transformed in the presence of ampicillin?
-The purpose is separation of those with the resistance gene and those without. - What will happen when bacterial cells that contain the pARA-R plasmid are not given arabinose?
-When they are without the arabinose the RFP gene will not be expressed and there will be no red glow. - In the lab, you will add samples of the control group P– and the treatment group P+ to plates that contain various combinations of Luria Broth (LB), ampicillin, and the sugar arabinose. Predict the growth on each dish.
-I predict that Plate I (LB) will have non-glowing growth on both sides. I predict that Plate II (LB/Amp) will have non-glowing growth on the P+ side. For Plate III (LB/Amp/Ara), I predict a glowing red colony.
During 5a:
- Look at the results of your transformation. Do your actual results match your predicted results? If not, what differences do you see, and what are some explanations for these differences?
-Our predictions were right except there was no red glowing on the LB/Amp/Ara plate. Glowing on that plate was the predicted result, however it did not occur. - How many red colonies were present on your LB/amp/ara plate?
-There were no red colonies that appeared on the plate. - Why did the red colonies only appear on the LB/Amp/Ara plate and not the LB/Amp plate?
-They need Ara-C in order for transcription to occur and gene expression. - Recombinant plasmids are engineered so that they can replicate in the cell independently of the chromosome replication. Why is it important to have multiple copies of a recombinant plasmid within a cell?
-It is important because there is extra for cell growth and in case of failure during transcription. - How is the information encoded in the RFP gene expressed as a trait? Be sure to use what you have previously learned about gene expression and the relationship between DNA, RNA, protein, and traits.
-The expression happen when the DNA is transcribed into protein using the central dogma DNA->mRNA->Protein. - Why is it possible for bacteria to make a human protein, such as insulin, or a sea anemone protein, such as the red fluorescent dye?
-All genes have the same central dogma (DNA -> mRNA ->Protein), so they can be expressed in seemingly different organisms. (Lab 2a: Question 4)
Before 6:
- How can solutions of different salt concentrations, which will unfold proteins to varying degrees, be used to help purify red fluorescent protein using column chromatography?
-Proteins are unfolded in the high salt concentrated buffer. The proteins that don't flow out the coulmn are folded again when lower salt concentrations are added, so they leave the column.
During 6:
- Why is a protein’s conformation important for carrying out its function?
-It determines the promoter regions of each protein. These different regions give each protein different functions. - What properties of the amino acids in a protein relate to protein folding?
-The sequence of amino acids determines it folds into a protein. - Does the eluate containing your red fluorescent protein appear less bright or brighter than it did in the cell lysate following centrifugation? If there is a noticeable difference in the intensity of the red color, what might account for that?
-The RFP-containing eluate is brighter than the cell-lysate, because the eluate contains the most RFP,while the cell lysate contains all the cells. - What characteristic of red fluorescent protein is used as the basis for separation by column chromatography?
-The red florescent protein attached to the resin column when unfolded. - How might the column chromatography procedure be adjusted or modified to increase the purity of the red fluorescent protein sample?
-If we repeated the experiment or lab with more wash buffers, and were more careful about collection, we would increase the expression of the RFP and we would see the protein more.
ANALYSIS/CONCLUSION
During this RFP lab, we tried to make a red bacteria colony by inserting genes. While the bacteria colony did not glow red, we still succeeded in part because the bacteria still had the red fluorescent protein gene. We isolated the RFP gene using restriction enzymes, and then double checked we had the gene with gel electrophoresis. After that, we injected the gene into the bacteria and grew the colony on a petri dish. The last step was using recombinant plasmids and chromatography to separate the protein into a tube.
Reflection
My group worked very well together during this project. We shared the workload evenly, explained confusing parts of the lab to other group members, and worked through the complicated procedure together. My lab skills increased, as well as my understanding of biotechnology and DNA structure. Overall, this was the best ;ab so far in terms of productivity and learned material.